Porcine Reproductive and Respiratory Syndrome
PRRSv biosecurity – key points to
stop entry to your farm
PRRSv biosecurity – reducing the spread
around your farm
Other names
|
PRRS, Blue Ear
Disease, Mystery Swine Disease, Swine Infertility and Respiratory Syndrome
(SIRS), Porcine Endemic Abortion and Respiratory Syndrome (PEARS) |
|||
|
Causal agent |
Virus. Porcine
Reproductive and Respiratory Syndrome Virus – RNA enveloped Differences in virulence are being detected through
variation in the structural genes. The
highly pathogenic form in China is associated with two deletions in NPS2
region. |
|||
|
Age group |
Adult: Clinical signs generally reproductive, mild fever
and anorexia |
|||
|
Piglets through to finishing: clinical signs generally associated
with secondary infections |
||||
|
Clinical signs |
||||
|
Naive herds |
Reproductive losses and a decreased farrowing rate |
|||
|
Early farrowings, at 105 to 112 days |
||||
|
Increase in stillborn, mummified and weak liveborn pigs |
||||
|
Increased pre-weaning mortality often associated with
increase in bacterial infections for example diarrhoea and greasy pig disease |
||||
|
Increased numbers of unthrifty pigs post weaning |
||||
|
Increased nursery mortality often associated with an increase
in bacterial infections for example post-weaning diarrhoea and meningitis |
||||
|
On established herds |
||||
|
Neonatal Pigs |
Respiratory Distress Unthrifty and failure to thrive
Increased secondary bacterial infections- diarrhoea and pneumonia |
|||
|
Growing pigs |
Increased mortality Decreased appetite Fever Rough hair
coat, unthrifty pigs |
|||
|
Increased respiratory problems, pneumonia and atrophic
rhinitis |
||||
|
Increased secondary bacterial infections for example meningitis,
Greasy pig disease |
||||
|
High Fever Problem in China associated with a novel
deletion of ORF 5 (NPS2) region |
||||
|
Adults |
North America strains can cause major reproductive
problems with massive abortions Late mummifications and abortions occur later in pregnancy
may be due to the fact that the early embryo and foetus have no receptors to
PRRSv and are therefore unaffected by the virus. But as they mature, they can become
infected. |
|||
|
|
|
|
|
|
|
Later mummified piglets with early farrowing |
Sick pig with complicated PRRSv |
“Blue ears” in a sow |
Abortions |
|
|
Infectivity |
||||
|
The virus particles have an envelope and rapidly becomes
inactivated in the environment and in the presence of disinfectants |
||||
|
Pig to pig contact is the major means of spread, through
infected faeces, urine and milk to piglets without colostral antibodies |
||||
|
Transmission through needles and insects is possible,
especially when blood transfer occurs |
||||
|
Air transmission possible, but mainly when major outbreaks
are present |
||||
|
While virus particles are seen in boar semen for 90+ days and
experimentally gilts can contract the disease through insemination. Thousands of inseminations from
serologically positive boars to naive herds has not resulted in the spread of
disease, therefore the risk through AI is small |
||||
|
When the disease first enters a country or new area, the
level of disease locally can be very high and aerosol spread possible. Once the disease has stabilised in an area
the risk of disease spread by semen or air is reduced. |
||||
Post-mortem
lesions
|
||||
|
There are very few visible post-mortem changes associated
with PRRSv, majority of the signs relate to secondary infections. Histologically the major finding is a
interstitial pneumonia and lack of air spaces. The
disease selectively kills the lung macrophage, essential for the defense of
the lung. The macrophages are killed
or damaged for 26 days. After 7 weeks
of age the alveolar macrophage becomes more resistant to PRRSv infection |
||||
|
|
|
|
|
|
Healthy macrophage
|
Dead macrophage |
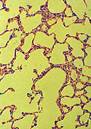
Normal lung |
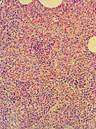
Interstitial pneumonia |
|
Diagnosis
|
||||
PRRSv
is suspected on the basis of the clinical signs
|
||||
|
The presence of PRRSv on a unit is confirmed by the use of
antibody tests. However, it can take 2-3 weeks for the antibody level to
rise before the test becomes positive.
Unfortunately the antibodies may also disappear 6 months after
exposure. |
||||
|
Examination of the lung tissue by histology
-immunohistochemistry can reveal the organism in the lung |
||||
|
PCR examination of tissues, in particular used for semen and
serum examination |
||||
|
Gene sequencing can be useful to monitor epidemiology of
PRRSv between farms |
||||
Treatment
and control
|
||||
|
Infected Herds |
There
is no specific anti viral treatment for PRRSv infection
|
|||
|
The
treatment regimes aim to minimise the effect of secondary infections. Aim to keep the pigs warm and in the
draught free environment and possibly increase feed density to compensate for
the anorexia. Review
the control measures for the secondary infections. Elimination of
PRRSv from units |
||||
|
Control |
SEW
programmes can help to control the spread of the disease around the farm and
minimise the effect of the disease on the farm's economy |
|||
|
All-in/
all-out and hygiene are essential precursors to controlling the disease |
||||
|
Current
live vaccines result in excretion from vaccinated pigs and therefore cannot
be used on PRRSv negative herds. The
use of live vaccines in incoming breeding animals in PRRS +ve herds helps to
maintain farm stability. The
vaccinated stock must be kept separate from the farm until shedding has
stopped |
||||
|
Home
(Autogenous) vaccines from serum or tonsilar scrape therapy may be utilised
to help gilt and boar introduction programmes. These should be restricted to the single
farm |
||||
|
Gilts
and boars must be stabilised before service. Discuss introduction programmes with your
veterinarian |
||||
|
Vaccines |
Dead
vaccines generally confer little or no protection
in naive animals, but it will reduce excretion of virus and assist reducing
farm clinical signs in infected herds. Modified
live vaccines (MLV) – these can be very variable in response
depending on the modification carried out.
Several MLV can cause severe clinical signs without field virus. In addition, there can be little protection
provided for heterologous virus strains.
Allowing sufficient time between vaccination and field infection
essential part of control. MLV
general reduce excretion of virus particles. |
|||
|
Review
fly and mosquito control programmes |
||||
|
PRRS-ve herds |
Before
purchasing breeding or other incoming stock ensure you match serostatus. Unfortunately the testing
procedures are not 100% accurate.
Practice on-farm AI collection, do not rely on a commercial AI stud |
|||
Common
differentials
|
||||
|
The clinical signs associated with Swine Influenza
can mimic many of the signs of PRRSv |
||||
|
Zoonotic implications |
||||
|
There are no zoonotic implications |
||||
Other issues
relating to Porcine Reproductive and Respiratory Syndrome Virus
PRRSv biosecurity – key points to
stop entry to your farm
PRRSv biosecurity – reducing the
spread around your farm
Treatment and control following a PRRSv break in a naïve
herd
Eradication of PRRSv in a positive
herd without depopulation
Biosecurity for
PRRSv and Biosecurity in
detail
PRRSv the genome and genetic identity
|
Porcine Reproductive and Respiratory Syndrome Virus |
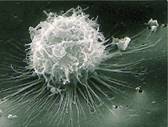
None enveloped, Positive sense single stranded RNA virus of the family Arteriviridae. The Arteriviridae and Coronaviridae families are combined into the Nidovirales order |
|
|
The appearance down the electron microscope. Dr
KJ Yoon |
|
|
|
The general layout of the
15 kb of the PRRSv genome, illustrating the two long Open Reading Frame
(ORF)1 and ORF2 and the smaller other frames.
Note there are some overlapping with the ORF |
There are 7 regions of the positive sense RNA genome of the PRRSv genome.
|
ORF 1 |
RNA replicase ORF1a and
ORF1b |
None structural proteins |
|
ORF 2 |
Minor membrane glycoprotein
GP2a GP2b |
Structural proteins |
|
ORF 2-7 |
Nucleocapsid protein N,
nucleolar localization |
|
|
ORF 3 |
Membrane glycoprotein GP3 |
|
|
ORF 4 |
Membrane glycoprotein GP4 |
|
|
ORF 5 |
Major membrane glycoprotein
GP5 |
|
|
ORF 6 |
Membrane associated protein
- M |
|
|
ORF 7 |
Nucleocapsid protein - N |
The GP5 protein is the most variable structural
protein, with only 51-55% aminoacid identity between North American and
European isolates, whereas the M protein is the most conserved protein with
78-80% aminoacid identity.`
Identification of the PRRSv genome is through two systems:
Restriction pattern
using MLU1; SacII and HincII enzymes
For example the predicted RFLP Pattern of the PRRSv isolated illustrated below is a
1-? - 4 Atypical Hinc II Pattern. This method does not produce satisfactory results.
Sequence bases within
ORF5
The ORF5 area is selected because this area demonstrates a high degree of variability and evolutionary change.
This results in a table of results – for example – chronograph
A=Adenine; T= Thymine – Uracil in original RNA; G=Guanine; C=Cytosine as the base pairs. Each 3 base pairs (a codon) codes for a different aminoacid, which when combined together produce the protein. Note the RNA is converted into a cDNA for sequencing as the RNA degenerates too quickly. However, the pattern of bases represents the actual Codons for their respective aminoacids.
|
10 |
20 |
30 |
40 |
Bases |
||||||||||||
|
atg |
ttg |
ggg |
aaa |
tgc |
ttg |
acc |
gcg |
ggc |
tgc |
tgc |
tcg |
caa |
ttg |
ctt |
ttt |
48 |
|
ttg |
tgg |
tgt |
atc |
gtg |
ccg |
ttc |
tgt |
ttt |
gtt |
gcg |
ctc |
gtc |
aac |
gcc |
gac |
96 |
|
AAC |
aac |
agc |
agc |
tcc |
cat |
tta |
cag |
ttg |
att |
tat |
aac |
ctg |
aca |
ata |
tgt |
144 |
|
gag |
ctg |
aat |
ggc |
aca |
gat |
tgg |
cta |
act |
aca |
aat |
ttt |
gat |
tgg |
gca |
gtg |
192 |
|
gag |
acc |
ttt |
gtc |
atc |
ttt |
cct |
gta |
ttg |
act |
cac |
atc |
gtc |
tcc |
tat |
ggt |
240 |
|
Gcc |
ctc |
acc |
acc |
agc |
cat |
ttc |
ctt |
gac |
aca |
gtc |
ggt |
ttg |
gtc |
act |
gtg |
288 |
|
tcc |
gcc |
gcc |
gga |
tac |
tgc |
cac |
ggg |
cgg |
tat |
gtc |
cta |
agt |
agc |
att |
gtg |
336 |
|
gct |
gtc |
tgc |
gcc |
ctg |
gcc |
gcg |
ctg |
att |
tgc |
ttc |
gcc |
atc |
agg |
ctg |
acg |
384 |
|
aaa |
aac |
tgc |
atg |
tcc |
tgg |
cgc |
tac |
tca |
tgt |
act |
aga |
tat |
act |
aac |
ttt |
432 |
|
ctt |
cta |
gac |
acc |
aag |
ggg |
aaa |
ctc |
tat |
cgt |
tgg |
cgg |
tct |
ccc |
gtc |
atc |
480 |
|
ata |
gag |
aaa |
ggg |
gga |
aaa |
atc |
gag |
gtt |
aac |
ggt |
cac |
ttg |
atc |
gac |
ctc |
528 |
|
aag |
aga |
gtt |
gtc |
ctt |
gat |
ggt |
tcc |
gcg |
gca |
act |
cct |
gta |
acc |
aaa |
gtt |
576 |
|
tcaa |
gcg |
gaa |
caa |
tgg |
tgt |
cgt |
cct |
tag |
603 |
This can then be compared with other published sequences, from vaccine strains for example, or from other sequenced isolated from the farm or possible source. When there is less than 99% difference (<6 nucleotides), the isolates can be considered similar or at least related.
For example, the above isolate was classified as similar to the following sequences:
|
Ingelvac ATP |
PrimePac |
RespPRRS |
Suvaxyn |
PRRomiSe |
Lelystad |
|
86.1% |
86.2% |
86.7% |
86.7% |
86.9% |
53.4% |
Therefore, this isolate was unrelated to any vaccine strain and the European strain (Lelystad) of PRRSv.
These patterns do not infer anything about virulence. Also note that the technique examines only 4% of the total genome of the virus.
Also remember that it takes 3 base pairs to code for an aminoacid and several types of codons code for the same aminoacid – so a change in a single base may not necessarily result in a change in aminoacid selected.